TREEPEACE
FROM HOLOCENE TO ANTHROPOCENE: THE PACE OF MICROEVOLUTION IN TREES
About TREEPEACE
The TREEPEACE project is funded by the European Research Council.
Project duration is 6 years, from 2014 to 2020.
The principal investigator of the project is Dr. Antoine Kremer and the host institution is the French National Institute of Agronomy Research INRA, UMR BIOGECO (BIODIVERSITY GENES & COMMUNITIES), in Bordeaux, France.
The research leading to these results has received funding from the European Research Council under the European Union’s Seventh Framework Programme (FP/2007-2013) / ERC Grant Agreement n. 339728
Summary
There are widespread concerns that trees, due to their long life-span, are not able to cope with the rapid ongoing climate change. While many studies have so far investigated potential impacts of climate change on forests, much less attention has been given to the potential evolutionary responses of tree populations. However there is a large body of evidence stemming from experimental evolutionary genetics showing that adaptive differentiation has extensively occurred during past environmental changes. This project challenges these views and explores the pace at which evolutionary change has taken place during past gradual and under current rapid environmental change. It builds on the reconstruction of evolutionary trajectories during the Holocene and Anthropocene to infer evolutionary rates, taking temperate oaks as a case study. The project assembles insights and contributions from paleobotany, ecology, ecophysiology, genetics, genomics and evolution in a generic framework for the assessment and prediction of rates of evolution at different hierarchical levels (genome to phenome). Beyond the assessments of past and current evolutionary change, the project provides an integrative simulation framework that will allow the monitoring and prediction of adaptive responses of trees under various evolutionary scenarios.

Provenances and progeny test during autumn 2015, INRA Pierroton, Cestas, France. © Laura Truffaut
Introduction
How much do tree population evolve? This is a central question for population biologists, but also a crucial issue nowadays for forest managers (Lindner et al., 2010; Lugo, 2015). There are indeed concerns that trees may not be able to cope with climate change due to their very long generation turnover. The likely decoupling between climate change velocity and the rate of evolutionary change has triggered recent research both at the theoretical and experimental level (Loarie et al., 2009). These concerns are even more acute in very long lived tree species as oaks (Kremer, 2013). Exploring the pace at which evolutionary change has taken place in sessile oak (Quercus petraea, a widespread European white oak) during past gradual and under current rapid environmental change, is the main focus of the Treepeace project (http://localhost/treepeace/), that was recently granted by the European Union as an ERC project (European research Council). This project builds on the reconstruction of evolutionary trajectories during the Holocene and Anthropocene and assembles insights from paleobotany, ecology, ecophysiology, genetics, genomics and evolution to track and monitor past and ongoing genetic changes during well documented historical warming periods: the postglacial period, the period following the little ice age and contemporary times. Part of the investigations are based on fossil remains, while analysis on extant –but very old- stands are used to document phenotypic or molecular changes over shorter time scales. Traditional approaches aiming at assessing evolutionary change adopted usually synchronic comparisons of extant provenances or populations originating from contrasting ecological sites. Their observed differences under common garden conditions were then interpreted as the results of different evolutionary forces. However such inferences allow only to estimate divergence rates, which are only a lower bound of the real evolutionary rates (Hendry and Kinnisson, 1999). In the Treepeace project we adopted an allochronic approach where the same populations are monitored at different points of time, over successive generations. While the project has started two years ago, very few results are available at this stage. The present article will therefore only provide an overview of the project and the methodological approaches used. We hope to offer at the next IOS meeting convincing results that oaks indeed are able to evolve substantially within a limited number of generations.
Working hypothesis and rationale for choosing oaks as a study material
This project falls within the pivotal question of population adaptive responses to ongoing climate change. If we are to evaluate future tree responses to climatic and anthropogenic changes, it is of utmost importance to learn from tree evolutionary trajectories in response to past changes. The need for retrospective investigations is particularly important given the growing body of evidence from various sources (paleobotany, population genetics, genetic surveys) that trees responded rapidly to post-glacial warming (Kremer et al., 2010). The rationale for using oaks as model species are based on the following background results and facts:
- There is experimental evidence based on common garden experiments of large genetic differentiation of adaptive traits, that has accumulated during the Holocene (Kremer et al., 2010). These observations have recently been confirmed at the genome level along altitudinal gradients mimicking temperature changes (Alberto et al., 2013).
- There is theoretical and experimental support that evolutionary changes are widespread in the genome, but that the amount of change at each genomic location is extremely small (Le Corre and Kremer, 2012). Thus a genome-wide exploration is needed to identify the genomic targets of evolution.
- The evolutionary trajectories and migration pathway of oaks during the Holocene have been reconstructed at a fine spatial grain level, by using a combination of phylogeographic investigations on extant populations and paleobotanical investigations (Petit et al., 2002).
- Oak macrofossils can be numerous and abundant and can be compiled from different sources for DNA extraction. They are often well-dated and well-described by dendrochronological and archeological means.
- A reference genome of Quercus robur, the most widespread oak species in Europe has recently been obtained (Plomion et al., 2015). Earlier experiments for retrieving DNA from wood samples (Deguilloux et al. 2006) together with recent progress in ancient DNA extraction, library preparation and genetic capture approaches (Gansauge et al. 2014) should allow to investigate allochronic evolutionary changes in the oak genome.
This brief review provides the foreground arguments to the working hypothesis of the Treepreace project. This hypothesis contends that the very wide genetic diversity that resides within tree species, and particularly oaks, might be the driving force of substantial genetic change in one single generation. In other words genetic diversity, which is the fuel of evolution, might compensate for the long generation for shifting populations to cope with environmental change.
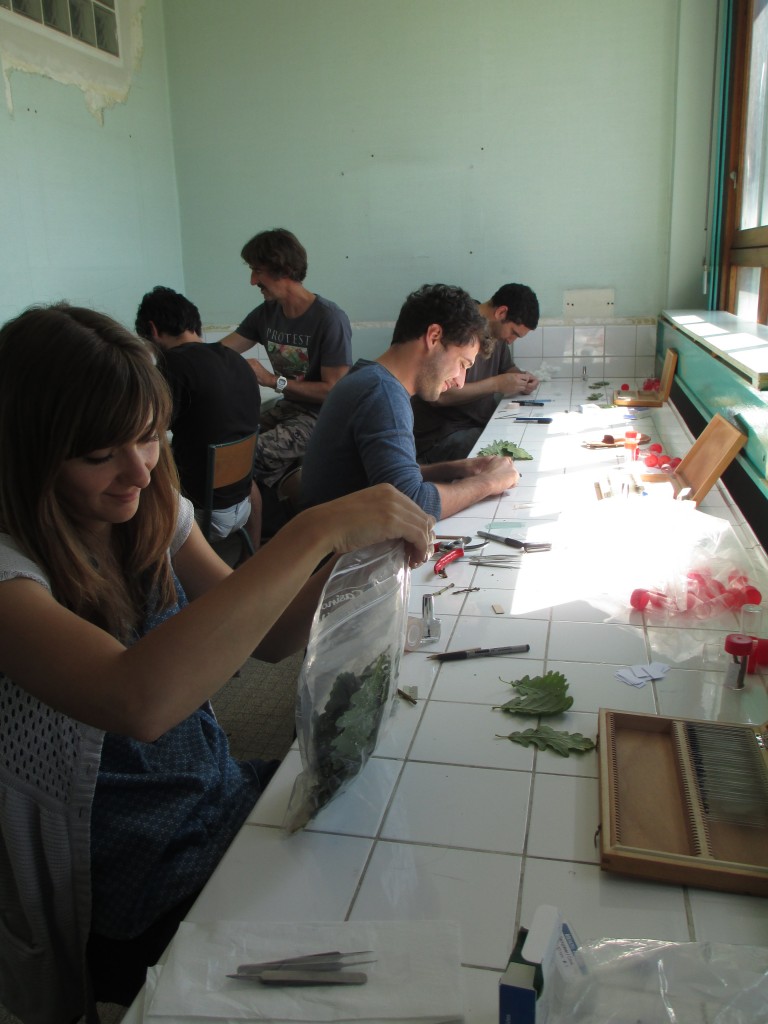

Leaf traits measurement during summer 2015, University of Bordeaux, Talence, France. © José M.Torres.
Assessing evolutionary change since the last glacial period
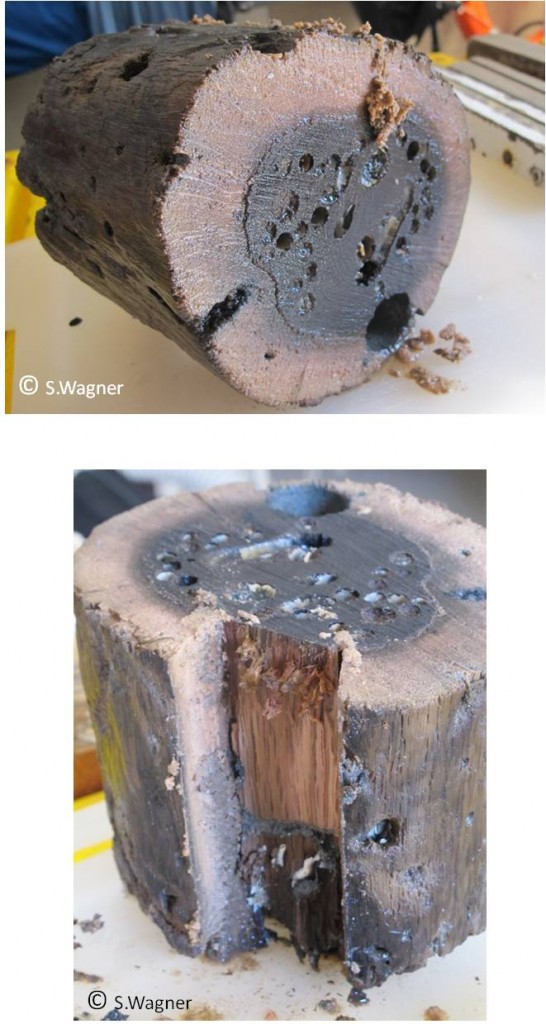
The Lateglacial and the Holocene (since 15000 years ago) are characterized by multiple short-term and long-term climate changes including a particularly steep temperature increase at the beginning of the Holocene (ca. 11,700 years ago) and a climate optimum reached during the mid-Holocene (Wright et al., 1993). The direct method to estimate evolutionary rate of European oaks during the Holocene is to compare the genetic diversity pattern of present-day populations to that of oak fossils collected in the ice age refugia. A first attempt to this end, was the recovery of aDNA from archaeological oak wood remains as old as 2,000 years (included in alluvial deposits and preserved in clay or sand) and the demonstration that small DNA fragment (53-87bp) could be amplified and cpDNA haplotypes obtained from this aDNA (Deguilloux et al. 2006). Although very scarce, other studies have shown that DNA information could be obtained from plant fossils preserved in water-influenced conditions (Manen et al. 2003, Pollmann et al. 2005), suggesting that anaerobic conditions could allow the preservation of DNA in submerged plant remains or that the rapid cementation around fossil tissues could form concretionary closed system before degradation is complete (Schweitzer 2004). We therefore preferentially collected oak samples to retrieve ancient DNA in remains that were conserved under water. So far we built a collection of about 400 oak woods coming from about 29 different sites from across Europe and from different temporal (700-10000 years) and environmental contexts (lacustrine, marine, terrestrial). The material was recovered from archeological excavations (piles from pile dwellings, planks from water well constructions) or anthropogenically uninfluenced natural contexts (submerged forests, burried forests) with the invaluable help of dendrochronologists and archeologists from across Europe (Figure 1). A prescreening of a subset of samples allowed to recover DNA from the collected wood samples. The majority of prescreened samples resulted in successful amplification in a qPCR experiment using a small chloroplast fragment. Checking pure extracts together with dilutions (1/10) revealed that inhibitory substances DNA could largely be removed during the extraction procedure. We also constructed a first genomic library of one of the samples (c. 6000 years) at the Centre of GeoGenetics at the University of Copenhagen (Denmark) in collaboration with Ludovic Orlando. Size distribution of the fragments of the library corresponded to the expectation for a typical DNA sample (primarily small fragments). There was sufficient DNA to send this library for sequencing (currently ongoing at the Centre of GeoGenetics).
|
Figure 1. Stem section from a watterlogged pile used for ancient DNA extraction. Earlier experiments and preliminary analysis of our own samples indicate that DNA was better preserved in wood samples that were conserved under water. Shown here is a section of an oak wood stem collected in Etang de Thau (near the city of Sète in the south of France), that is 3000 years old. One can easily recognize the sapwood, from which DNA is extracted. Waterlogged wood samples do exist either as subfossil remains of submerged stems in marine or lacustrine waters, or as piles used for human constructions. Pile dwellings (or stilt houses) do exist in many alpine lakes, where the first farmers established their settlements some 7000 years ago. More than 1000 settlements sites have been discovered and about 100 were recently classified as UNESCO World Heritage sites. |
Assessing evolutionary change since the Little Ice Age
The “Little Ice Age” (LIA) was a period that lasted loosely from the sixteenth to the nineteenth centuries, during which the climate was much cooler than under the medieval warm period (Fagan, 2000). Although there were regional differences throughout Europe, it is well accepted that temperatures were significantly lower between 1570 and 1900 in comparison to today’s climate (Büntgen and Hellman, 2014). More specifically periods like the Maunder minimum (1635 to 1665) and the early nineteenth century are characterized by reduced solar activity and increased volcanic activity (Crowley, 2000) and were among the colder periods during the LIA. The former period overlaps partly with the 30 years wars in Europe, which was also well known for his harsh climate (Büntgen et al., 2013). Incidentally, the warming period following the LIA coincides with the Anthropocene, that started during the industrial revolution when human impacts has driven the planet into a new geologic epoch (Crutzen and Stoemer, 2000). Within Treepeace, we attempt to explore whether there has been measurable evolutionary change either at the phenotypic or genomic level in oak forest during the warming period that elapsed since the LIA. We adopted an allochronic approach based on living trees, in contrast to the methodology employed for monitoring evolutionary change during the Holocene. The oldest sessile oaks forests in France stem from the mid 17th century, when Jean Baptiste Colbert, prime minister of the King Louis XIV, decreed that oak forests should be restored to produce timber for ship building (Devèze, 1962). One of the measures that were implemented by Colbert’s “ordonnance” was the transformation of oak coppice into high forest. Under even aged high forest management regimes, generation turnover is done via natural regeneration that produces very high densities of seedlings that may reach up to 1 million seedlings/ha in the case of oaks. Natural selection will reduce densities very rapidly within the first 10 years to approximately 50 000. Later selections are achieved by human interventions at systematic or selective cleanings and thinning ending finally to densities of about 100 trees per ha. Clearly under such cultural regimes, natural selection is still the main driving force of oak stands, as 95% of the seedlings are eliminated during the very early 10 to 15 years. These figures suggest therefore that today’s oldest oak stands in France resulted mainly from natural selection that occurred during the LIA. With the help of the French Forest Service (ONF, Office National des Forêts), we made an inventory of existing pure sessile oak forests comprising stands that originated during the LIA, and that met in addition the following criteria:
- Documentary and historical records of natural regeneration
- There is no chloroplast DNA (CpDNA) diversity within the stand. CpDNA fixation is a genetic indication that todays’ stands originated from repeated natural regenerations.
- No co-occurrence of other white oaks likely to cross with Sessile oak
This procedure resulted finally in the selection of three sessile oak forests: Bercé, Tronçais and RénoValdieu (Figure 2). Within each forest we identified 4 cohorts:
- Old cohort: more than 350 years
- Medium age cohort : between 150 and 180 years
- Young cohort: 50 years
- Juvenile cohort: less than 10 years.
The objective of the project is to assess genetic differentiation between the 4 cohorts of a given forest. We make the assumption that genetic differentiation is likely to result from different selective pressures that shaped the 4 cohorts during their juvenile phase. Implicitly, this assumption contends that no other source of differentiation has generated any noise to the differentiation. Such noise may for example originate from different “genetic histories” of the cohorts, or admixture with other white oak species.
In order to assess genetic differentiation between the cohorts, open pollinated progenies were collected in each cohort and in each forest and are currently raised in a nursery under common garden conditions. This material will ultimately be outplanted in the field for monitoring differentiation among the cohorts over the years. Leaf and cambium samples were collected on adults trees of the different cohorts for whole genome sequencing of all the cohorts in order to assess whether patterns of genomic differentiation can also be detected.
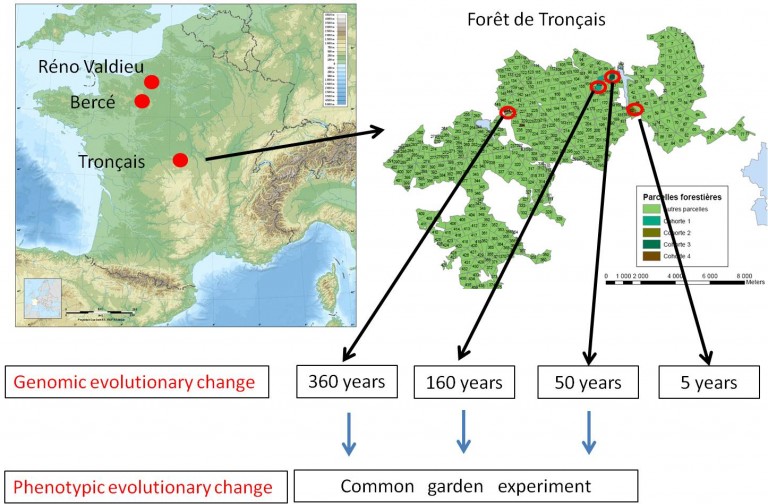
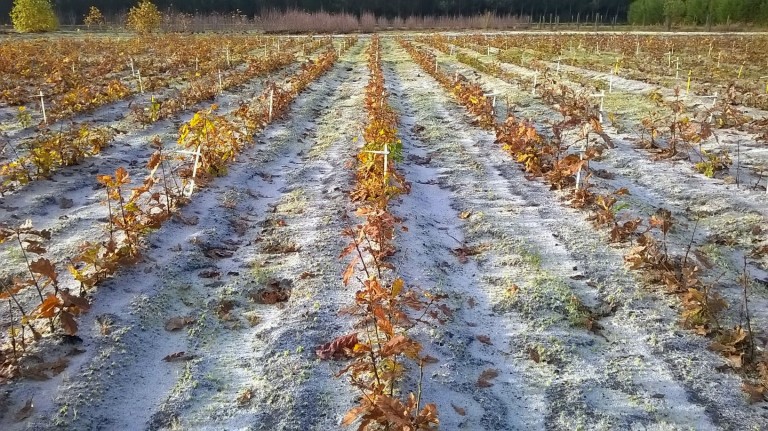
Figure 2. Sampling design for assessing evolutionary change since the Little Ice Age
Three pure sessile oak forests (Reno Valdieu, Bercé and Tronçais) comprising very old compartments originating from the mid seventeen century were selected based on documentary information. The dating was confirmed by tree ring analysis of a few felled trees. These three forest are managed as high forests under even aged regimes, as illustrated here in the case of Tronçais. The forest is subdivided in many contiguous compartments. The size of a compartment is between 15 to 30 ha and comprises trees belonging to the same age class. We selected 4 compartments corresponding to 4 cohorts of different ages: 360, 160, 50 and 5 years. About 50 trees were randomly sampled within each cohort, and open pollinated progenies were collected on 30 trees par cohort (except in the youngest cohort). Whole genome sequencing will be achieved for each cohort and phenotypic comparisons will be made in the common garden experiment to assess genetic differentiation between the cohorts. The same procedure is repeated in all three forests.
Assessing evolutionary change at contemporary time scales
This part of the project intends to estimate evolutionary changes over one single generation. Again allochronic methods which would allow to assess the same trait (at the same age) in two successive generation may be hardly possible due to obvious time constraints in longlived species as oaks. However under quantitative genetic principles, evolutionary change from one generation to the next under some form of selection can be predicted based on genetic parameters of the population, some of which can be estimated within one generation. Under standard quantitative genetic principles the evolutionary change ΔG between generation t and t+1 can be predicted at generation t even though the next generation has not been obtained (Lande and Arnold 1983):
ΔG=VG β (1)
Where VG is the genetic variance in the population and β is the selection gradient. Intuitively speaking, the selection gradient is a synthetic measure of the strength and direction of selection. In its mathematical formulation it is the regression coefficient, β, between the fitness of the tree, W, and the trait value, Z: β = cov (W,Z)/VZ (Lande and Arnold 1983). The project therefore aims at estimating VG and β in natura for a set of phenotypic traits likely to be adaptive under the current climatic change.
For this purpose we selected the Petite Charnie intensive study plot (ISP), which has been investigated over the past 30 years (Figure 3). Briefly an exhaustive genetic and phenotypic inventory has been conducted during the mid 90s on the 5 ha plot of the Petite Charnie ISP (Streiff et al.,1998,1999). At that time the ISP was a hundred year old mixed oak stand (Quercus petraea and Quercus robur) composed of 426 trees. All trees were fingerprinted, and grafted in an archive conservation plot. Detailed assessments of phenotypic attributes (growth, form, architecture, secondary metabolites, wood density ..) were made. A seed cutting was done in the mid 90s followed by a clear cutting of the remaining trees in 2000, when natural seeding was completed. A very dense regeneration developed after the clear cut. In the summer of 2014, when the regeneration was 15 to 20 years old, and after natural selection already reduced the initial densities, we systematically sampled 2500 saplings in the regeneration. By using genomic fingerprints in the juvenile and adult cohorts we were able to calculate the reproductive success of each parent tree, eg the number of offspring that a parent tree produced and that survived the initial natural selection (during the past 15 to 20 years). The reproductive success is a proxy of the fitness (W). The next steps will consist in comparing the fitness and the phenotypic trait value (Z) of each parent tree. Any correlation among these measures will indicate that the trait value is changed by natural selection. If that trait exhibits genetic variation (VG), eg the trait is heritable, then the change will be transmitted to the next generation. Heritability of the traits will also be estimated in natura (in the Petite Charnie ISP) by using standard quantitative genetic approaches.
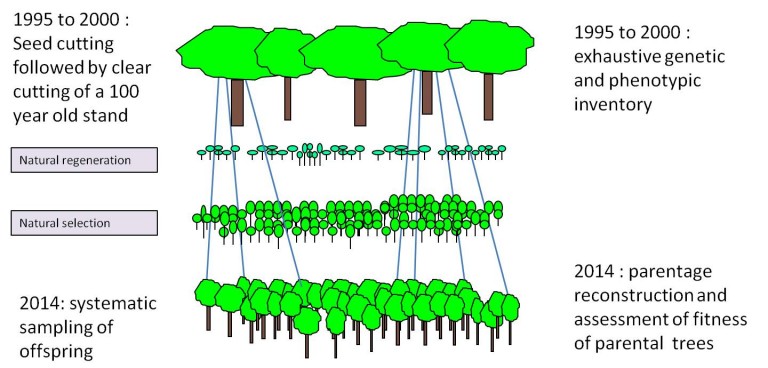
Figure 3. Assessing evolutionary change between two successive generations
This experiments aims at identifying phenotypic traits that are likely to undergo selection due to current climate change and to estimate how much genetic change can be predicted. Whether a trait is under selection depends if it is correlated to the fitness of a tree (see text). Here we aim at measuring the reproductive success of a tree as a proxy of fitness. The research is conducted in the Petite Charnie intensive study plot, where large scale genetic investigations have been conducted since 30 years in a mixed oak stand (Quercus petraea and Quercus robur). The stand was clear cut in 2000, when continuous natural regeneration was obtained. Fourteen years later, after strong natural selection occurred, we performed a parentage analysis using molecular fingerprints of the parental trees and the regeneration saplings. Parentage reconstruction permitted to estimate the reproductive success, eg the number of offspring produced by each parent. If trees that produce many offspring that survived the early stages, are for example early flushing, then the next generation will be shifted towards earlyness of flushing.
Conclusion
This Treepeace project aims to estimate the pace at which microevolution in oaks has and is taking place. The project will benefit from contributions and insights from paleobotany, archaeology, ecology, genomics and evolution in a generic framework for assessing and predicting rates of evolution. The time has come where these complementary approaches can be assembled and help to understand the complex evolutionary trajectories that allowed oaks to overcome past environmental crisis. There are of course challenging issues, as the retrieving of ancient DNA from the last glacial period, or the assessment of fitness of adult trees. But recent progress in the field of genomics of ancient DNA and evolutionary quantitative genetics should allow substantial progress in evolutionary biology of long lived species. We hope to be able to report of the expected achievements during the next meetings of the IOS.
Acknowledgements
I thank the European Union for the financial support of the research through the ERC project Treepeace. I am grateful to the numerous colleagues archaeologists and dendrochronologist that shared their experience and knowledge about ancient oak logs and trees, and allowed to access their archeological or fossil remains. I offer my warmest thanks to the staff of BIOGECO, doctorate students and postdotorates that contribute to the different activities of the Treepeace project.
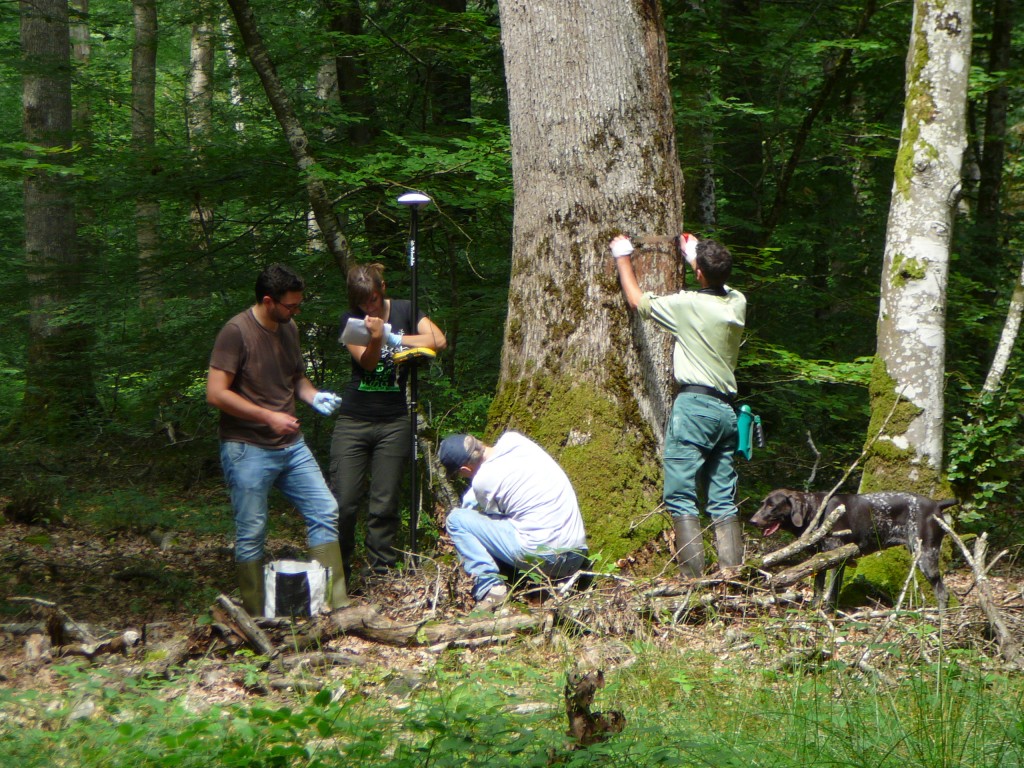

Cambium sampling in Tronçais Forest, summer 2014, Tronçais, France. © Didier Bert
References
Büntgen, U., and L. Hellmann. 2014. The little ice age in scientific perspective: cold spells and caveats. Journal of Interdisciplinary History 44: 353–368.
Büntgen, U., T. Kyncl, C. Ginzler, D.S. Jacks, J. Esper, W. Tegel, K.U. Heussner, and J. Kyncl. 2013. Filling the Eastern European gap in millennium-long temperature reconstructions. PNAS 110: 1773-1778
Crutzen, P. J., and E. F. Stoermer. 2000. « The ‘Anthropocene' ». Global Change Newsletter 41: 17–18.
Devèze, M.1962. La grande reformation des forêts royales sous Colbert (1661-1680). Annales de L’Ecole Nationale Des Eaux et Forêts 19 : 15-164
Fagan, B. 2000. The Little Ice Age. How climate made history. Basic Books, 235pages
Gansauge, M.T., and M. Meyer. 2014. Selective enrichment of damaged DNA molecules for ancient genome sequencing. Genome Research 24: 1543-1549.
Hendry, A.P, and M.T. Kinnison. 1999. Perspective: the pace of modern life: measuring rates of contemporary microevolution. Evolution 53 (6): 1637-1652
Kremer, A., V. Le Corre, R.J. Petit and A. Ducousso. 2010. Historical and contemporary dynamics of adaptive differentiation in European oaks. In : DeWoody A., Bickham J., Michler C., Nichols K., Rhodes G., Woeste K. (eds), p101-122 Molecular approaches in natural resource Conservation. Cambridge University Press
Kremer, A. 2013. Evolutionary responses of European oaks to climate change. International Oaks, 11-20
Le Corre, V., and A. Kremer. 2012. The genetic differentiation at quantitative trait loci under local adaptation. Molecular Ecology 21, 1548-1566
Lande, R., and S.J. Arnold. 1983. The measurement of selection on correlated characters. Evolution 37: 1210-1226
Lindner, M., M. Maroschek, S. Netherer, A. Kremer, A. Barbati, J. Garcia-Gonzalo, R. Seidl, S. Delzon, P. Corona, M. Kolström, M.J. Lexer, and M. Marchetti. 2010. Climate change impacts, adaptive capacity, and vulnerability of European forest ecosystems. Forest Ecology and Management 259 :698-709
Loarie, S.R., P.B. Duffy, H. Hamilton, G. Asner, C.B. Field, and D.D. Ackerly. 2009. The velocity of climate change. Nature 462: 1052-1055
Lugo, A.E., 2015. Forestry in the Anthropocene. Science 349:771
Manen J.F., L. Bouby, O. Dalnoki, P. Marinval, M. Turgay, and A. Schlumbaum. 2003. Microsatellites from archaeological Vitis vinifera seeds allow a tentative assignment of the geographical origin of ancient cultivars. Journal of Archaeological Science 30: 721e729.
Petit, R.J., S. Brewer, S. Bordacs, K. Burg, R. Cheddadi, E. Coart, J. Cottrell, U.M. Csaikl, B.C. van Dam, J.D. Deans, S. Espinel, S. Fineschi, R. Finkeldey, I. Glaz, P.G. Goiecochea, J.S. Jensen, A.O. König, A.J. Lowe, S.F. Madsen, G. Matyas, R.C. Munro, F. Popescu, D. Slade, H. Tabbener, S.M.G. de Vries, B. Ziegenhagen, J.-L. de Beaulieu, and A. Kremer. 2002. Identification of refugia and post-glacial colonisation routes of European white oaks based on chloroplast DNA and fossil pollen evidence. Forest Ecology and Management 156 (1-3):49-74.
Pollmann, B., S. Jacomet, and A. Schlumbaum. 2005. Morphological and genetic studies of waterlogged Prunus species from the Roman vicus Tasgetium (Eschenz, Switzerland), Journal of Archaeological Science 32: 1471e1480.
Schweitzer, M.H. 2004 Molecular paleontology: some current advances and problems, Annales de Paléontologie 90: 81e102
Streiff, R., T. Labbe, R. Bacilieri, H. Steinkellner, J. Glossl, and A. Kremer. 1998. Within-population genetic structure in Quercus robur L. and Quercus petraea (Matt.) Liebl. assessed with isozymes and microsatellites. Molecular Ecology 7: 317-328
Streiff, R., A. Ducousso, C. Lexer, H. Steinkellner, J. Gloessl, and A. Kremer. 1999. Pollen dispersal inferred from paternity analysis in a mixed oak stand of Quercus robur L. and Q. petraea (Matt.) Liebl. Molecular Ecology 8: 831-841.
Wright H.E., J.E. Kutzbach, T. Webb, W.F. Ruddiman, F.A. Street-Perrott, and P.J Bartlein. 1993. Global Climates since the Last Glacial Maximum, University of Minnesota Press, 584p

